 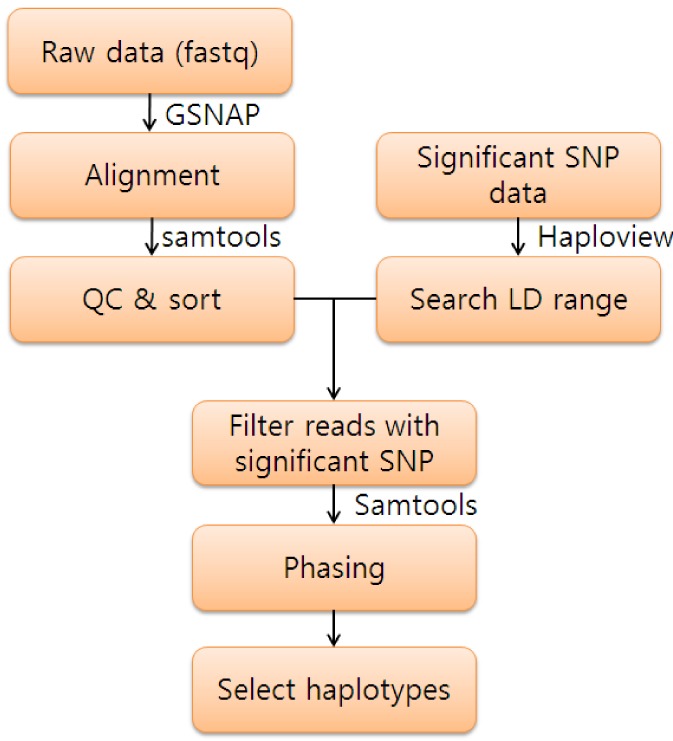
In the fifth field, a dot or a comma denotes a base identical to the reference a dot or a capital letter denotes a base from a read mapped on the forward strand, while a comma or a lowercase letter on the reverse strand. Each line consists of reference name, sorted coordinate, reference base, the number of reads covering the position and read bases. ( c) Simplified pileup output by SAMtools. Read r004 is aligned across an intron, indicated by the N operation. Preparing system Open a terminal by pressing Ctrl+Alt+T. For the installation tutorial of BCFtools, click here. We are going to install SAMtools only on Ubuntu. It consists of three separate packages: SAMtools, BCFtools, and HTSlib. The NM tag gives the number of mismatches. SAMtools is a software package for high-throughput sequencing data analysis 1. The hard clipping operation H indicates that the clipped sequence is not present in the sequence field. The last six bases of read r003 map to position 9, and the first five to position 29 on the reverse strand. Padding operations can be absent when an aligner does not support multiple sequence alignment. The CIGAR string for this alignment contains a P (padding) operation which correctly aligns the inserted sequences. The coordinate shown in SAM is the position of the first aligned base. 
Read r002 has three soft-clipped (unaligned) bases. According to FLAG 163 (=1 + 2 + 32 + 128), the read mapped to position 7 is the second read in the pair (128) and regarded as properly paired (1 + 2) its mate is mapped to 37 on the reverse strand (32). Notably, r001 is the name of a read pair. The ‘ line in the header section gives the order of reference sequences. ( a) Alignments of one pair of reads and three single-end reads. Example of extended CIGAR and the pileup output. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
